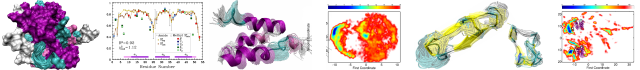Nucleic Acids Research
Interference activity of a minimal Type I CRISPR-Cas system from Shewanella putrefaciens
Type I CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)–Cas (CRISPR-associated) systems exist in bacterial and archaeal organisms and provide immunity against foreign DNA. The Cas protein content of the DNA interference complexes (termed Cascade) varies between different CRISPR-Cas subtypes. A minimal variant of the Type I-F system was identified in proteobacterial species including Shewanella putrefaciens CN-32. This variant lacks a large subunit (Csy1), Csy2 and Csy3 and contains two unclassified cas genes. The genome of S. putrefaciens CN-32 contains only five Cas proteins (Cas1, Cas3, Cas6f, Cas1821 and Cas1822) and a single CRISPR array with 81 spacers. RNA-Seq analyses revealed the transcription of this array and the maturation of crRNAs (CRISPR RNAs). Interference assays based on plasmid conjugation demonstrated that this CRISPR-Cas system is active in vivo and that activity is dependent on the recognition of the dinucleotide GG PAM (Protospacer Adjacent Motif) sequence and crRNA abundance. The deletion of cas1821 and cas1822 reduced the cellular crRNA pool. Recombinant Cas1821 was shown to form helical filaments bound to RNA molecules, which suggests its role as the Cascade backbone protein. A Cascade complex was isolated which contained multiple Cas1821 copies, Cas1822, Cas6f and mature crRNAs.
Structure and specificity of the RNA-guided endonuclease Cas9 during DNA interrogation, target binding and cleavage
CRISPR-associated endonuclease Cas9 cuts DNA at variable target sites designated by a Cas9-bound RNA molecule. Cas9's ability to be directed by single ‘guide RNA’ molecules to target nearly any sequence has been recently exploited for a number of emerging biological and medical applications. Therefore, understanding the nature of Cas9's off-target activity is of paramount importance for its practical use. Using atomic force microscopy (AFM), we directly resolve individual Cas9 and nuclease-inactive dCas9 proteins as they bind along engineered DNA substrates. High-resolution imaging allows us to determine their relative propensities to bind with different guide RNA variants to targeted or off-target sequences. Mapping the structural properties of Cas9 and dCas9 to their respective binding sites reveals a progressive conformational transformation at DNA sites with increasing sequence similarity to its target. With kinetic Monte Carlo (KMC) simulations, these results provide evidence of a ‘conformational gating’ mechanism driven by the interactions between the guide RNA and the 14th–17th nucleotide region of the targeted DNA, the stabilities of which we find correlate significantly with reported off-target cleavage rates. KMC simulations also reveal potential methodologies to engineer guide RNA sequences with improved specificity by considering the invasion of guide RNAs into targeted DNA duplex.
The Bacteroides sp. 3_1_23 Pif1 protein is a multifunctional helicase
ScPif1 DNA helicase is the prototypical member of a 5'-to-3' helicase superfamily conserved from bacteria to human and plays various roles in the maintenance of genomic homeostasis. While many studies have been performed with eukaryotic Pif1 helicases, including yeast and human Pif1 proteins, the potential functions and biochemical properties of prokaryotic Pif1 helicases remain largely unknown. Here, we report the expression, purification and biochemical analysis of Pif1 helicase from Bacteroides sp. 3_1_23 (BsPif1). BsPif1 binds to a large panel of DNA substrates and, in particular, efficiently unwinds partial duplex DNAs with 5'-overhang, fork-like substrates, D-loop and flap-like substrates, suggesting that BsPif1 may act at stalled DNA replication forks and enhance Okazaki fragment maturation. Like its eukaryotic homologues, BsPif1 resolves R-loop structures and unwinds DNA–RNA hybrids. Furthermore, BsPif1 efficiently unfolds G-quadruplexes and disrupts nucleoprotein complexes. Altogether, these results highlight that prokaryotic Pif1 helicases may resolve common issues that arise during DNA transactions. Interestingly, we found that BsPif1 is different from yeast Pif1, but resembles more human Pif1 with regard to substrate specificity, helicase activity and mode of action. These findings are discussed in the context of the possible functions of prokaryotic Pif1 helicases in vivo.
The rates of the major steps in the molecular mechanism of RNase H1-dependent antisense oligonucleotide induced degradation of RNA
Antisense oligonucleotides (ASOs) are most commonly designed to reduce targeted RNA via RNase H1-dependent degradation, however kinetic parameters for ASO-mediated targeting and subsequent cleavage and degradation of RNA in living cells are poorly understood. In this manuscript we use an inducible minigene system to determine the time course of ASO activity in the cell. Estimates of the time required for the ASO to enter and traverse the cell, scan the target mRNA, bind the cognate site, recruit RNase H1 and initiate cleavage, are presented in the context of transcription and mRNA processing rates. Data are also presented which indicate that rates for RNase H1-dependent ASO-mediated degradation of the targeted RNAs are different for nuclear-retained versus RNAs exported to the cytoplasm and that the level of RNase H1 in the cell and cellular compartments is limiting to the rate of ASO activity. In both cellular compartments RNase H1 ASOs essentially double the endogenous rates of clearance of the target RNA. Overexpression of Escherichia coli RNase H1 or the presence of multiple cognate sites each further increase the rate of target RNA degradation.
Global analysis of RNA cleavage by 5'-hydroxyl RNA sequencing
RNA cleavage by some endoribonucleases and self-cleaving ribozymes produces RNA fragments with 5'-hydroxyl (5'-OH) and 2',3'-cyclic phosphate termini. To identify 5'-OH RNA fragments produced by these cleavage events, we exploited the unique ligation mechanism of Escherichia coli RtcB RNA ligase to attach an oligonucleotide linker to RNAs with 5'-OH termini, followed by steps for library construction and analysis by massively parallel DNA sequencing. We applied the method to RNA from budding yeast and captured known 5'-OH fragments produced by tRNA Splicing Endonuclease (SEN) during processing of intron-containing pre-tRNAs and by Ire1 cleavage of HAC1 mRNA following induction of the unfolded protein response (UPR). We identified numerous novel 5'-OH fragments derived from mRNAs: some 5'-OH mRNA fragments were derived from single, localized cleavages, while others were likely produced by multiple, distributed cleavages. Many 5'-OH fragments derived from mRNAs were produced upstream of codons for highly electrostatic peptides, suggesting that the fragments may be generated by co-translational mRNA decay. Several 5'-OH RNA fragments accumulated during the induction of the UPR, some of which share a common sequence motif that may direct cleavage of these mRNAs. This method enables specific capture of 5'-OH termini and complements existing methods for identifying RNAs with 2',3'-cyclic phosphate termini.
Tailor: a computational framework for detecting non-templated tailing of small silencing RNAs
Small silencing RNAs, including microRNAs, endogenous small interfering RNAs (endo-siRNAs) and Piwi-interacting RNAs (piRNAs), have been shown to play important roles in fine-tuning gene expression, defending virus and controlling transposons. Loss of small silencing RNAs or components in their pathways often leads to severe developmental defects, including lethality and sterility. Recently, non-templated addition of nucleotides to the 3' end, namely tailing, was found to associate with the processing and stability of small silencing RNAs. Next Generation Sequencing has made it possible to detect such modifications at nucleotide resolution in an unprecedented throughput. Unfortunately, detecting such events from millions of short reads confounded by sequencing errors and RNA editing is still a tricky problem. Here, we developed a computational framework, Tailor, driven by an efficient and accurate aligner specifically designed for capturing the tailing events directly from the alignments without extensive post-processing. The performance of Tailor was fully tested and compared favorably with other general-purpose aligners using both simulated and real datasets for tailing analysis. Moreover, to show the broad utility of Tailor, we used Tailor to reanalyze published datasets and revealed novel findings worth further experimental validation. The source code and the executable binaries are freely available at https://github.com/jhhung/Tailor.
Nucleotidyl transferase assisted DNA labeling with different click chemistries
Here, we present a simple, modular and efficient strategy that allows the 3'-terminal labeling of DNA, regardless of whether it has been chemically or enzymatically synthesized or isolated from natural sources. We first incorporate a range of modified nucleotides at the 3'-terminus, using terminal deoxynucleotidyl transferase. In the second step, we convert the incorporated nucleotides, using either of four highly efficient click chemistry-type reactions, namely copper-catalyzed azide-alkyne cycloaddition, strain-promoted azide-alkyne cycloaddition, Staudinger ligation or Diels-Alder reaction with inverse electron demand. Moreover, we create internal modifications, making use of either ligation or primer extension, after the nucleotidyl transferase step, prior to the click reaction. We further study the influence of linker variants on the reactivity of azides in different click reactions. We find that different click reactions exhibit distinct substrate preferences, a fact that is often overlooked, but should be considered when labeling oligonucleotides or other biomolecules with click chemistry. Finally, our findings allowed us to extend our previously published RNA labeling strategy to the use of a different copper-free click chemistry, namely the Staudinger ligation.
A novel hybrid single molecule approach reveals spontaneous DNA motion in the nucleosome
Structural dynamics of nucleic acid and protein is an important physical basis of their functions. These motions are often very difficult to synchronize and too fast to be clearly resolved with the currently available single molecule methods. Here we demonstrate a novel hybrid single molecule approach combining stochastic data analysis with fluorescence correlation that enables investigations of sub-ms unsynchronized structural dynamics of macromolecules. Based on the method, we report the first direct evidence of spontaneous DNA motions at the nucleosome termini. The nucleosome, comprising DNA and a histone core, is the fundamental packing unit of eukaryotic genes that must be accessed during various genome transactions. Spontaneous DNA opening at the nucleosome termini has long been hypothesized to enable gene access in the nucleosome, but has yet to be directly observed. Our approach reveals that DNA termini in the nucleosome open and close repeatedly at 0.1–1 ms–1. The kinetics depends on salt concentration and DNA–histone interactions but not much on DNA sequence, suggesting that this dynamics is universal and imposes the kinetic limit to gene access. These results clearly demonstrate that our method provides an efficient and robust means to investigate unsynchronized structural changes of DNA at a sub-ms time resolution.
A modular open platform for systematic functional studies under physiological conditions
Any profound comprehension of gene function requires detailed information about the subcellular localization, molecular interactions and spatio-temporal dynamics of gene products. We developed a multifunctional integrase (MIN) tag for rapid and versatile genome engineering that serves not only as a genetic entry site for the Bxb1 integrase but also as a novel epitope tag for standardized detection and precipitation. For the systematic study of epigenetic factors, including Dnmt1, Dnmt3a, Dnmt3b, Tet1, Tet2, Tet3 and Uhrf1, we generated MIN-tagged embryonic stem cell lines and created a toolbox of prefabricated modules that can be integrated via Bxb1-mediated recombination. We used these functional modules to study protein interactions and their spatio-temporal dynamics as well as gene expression and specific mutations during cellular differentiation and in response to external stimuli. Our genome engineering strategy provides a versatile open platform for efficient generation of multiple isogenic cell lines to study gene function under physiological conditions.
Disturbance-free rapid solution exchange for magnetic tweezers single-molecule studies
Single-molecule manipulation technologies have been extensively applied to studies of the structures and interactions of DNA and proteins. An important aspect of such studies is to obtain the dynamics of interactions; however the initial binding is often difficult to obtain due to large mechanical perturbation during solution introduction. Here, we report a simple disturbance-free rapid solution exchange method for magnetic tweezers single-molecule manipulation experiments, which is achieved by tethering the molecules inside microwells (typical dimensions–diameter (D): 40–50 μm, height (H): 100 μm; H:D~2:1). Our simulations and experiments show that the flow speed can be reduced by several orders of magnitude near the bottom of the microwells from that in the flow chamber, effectively eliminating the flow disturbance to molecules tethered in the microwells. We demonstrate a wide scope of applications of this method by measuring the force dependent DNA structural transitions in response to solution condition change, and polymerization dynamics of RecA on ssDNA/SSB-coated ssDNA/dsDNA of various tether lengths under constant forces, as well as the dynamics of vinculin binding to α-catenin at a constant force (< 5 pN) applied to the α-catenin protein.
MINT: software to identify motifs and short-range interactions in trajectories of nucleic acids
Structural biology experiments and structure prediction tools have provided many high-resolution three-dimensional structures of nucleic acids. Also, molecular dynamics force field parameters have been adapted to simulating charged and flexible nucleic acid structures on microsecond time scales. Therefore, we can generate the dynamics of DNA or RNA molecules, but we still lack adequate tools for the analysis of the resulting huge amounts of data. We present MINT (Motif Identifier for Nucleic acids Trajectory) — an automatic tool for analyzing three-dimensional structures of RNA and DNA, and their full-atom molecular dynamics trajectories or other conformation sets (e.g. X-ray or nuclear magnetic resonance-derived structures). For each RNA or DNA conformation MINT determines the hydrogen bonding network resolving the base pairing patterns, identifies secondary structure motifs (helices, junctions, loops, etc.) and pseudoknots. MINT also estimates the energy of stacking and phosphate anion-base interactions. For many conformations, as in a molecular dynamics trajectory, MINT provides averages of the above structural and energetic features and their evolution. We show MINT functionality based on all-atom explicit solvent molecular dynamics trajectory of the 30S ribosomal subunit.
ATM-dependent phosphorylation of MRE11 controls extent of resection during homology directed repair by signalling through Exonuclease 1
The MRE11/RAD50/NBS1 (MRN) complex plays a central role as a sensor of DNA double strand breaks (DSB) and is responsible for the efficient activation of ataxia-telangiectasia mutated (ATM) kinase. Once activated ATM in turn phosphorylates RAD50 and NBS1, important for cell cycle control, DNA repair and cell survival. We report here that MRE11 is also phosphorylated by ATM at S676 and S678 in response to agents that induce DNA DSB, is dependent on the presence of NBS1, and does not affect the association of members of the complex or ATM activation. A phosphosite mutant (MRE11S676AS678A) cell line showed decreased cell survival and increased chromosomal aberrations after radiation exposure indicating a defect in DNA repair. Use of GFP-based DNA repair reporter substrates in MRE11S676AS678A cells revealed a defect in homology directed repair (HDR) but single strand annealing was not affected. More detailed investigation revealed that MRE11S676AS678A cells resected DNA ends to a greater extent at sites undergoing HDR. Furthermore, while ATM-dependent phosphorylation of Kap1 and SMC1 was normal in MRE11S676AS678A cells, there was no phosphorylation of Exonuclease 1 consistent with the defect in HDR. These results describe a novel role for ATM-dependent phosphorylation of MRE11 in limiting the extent of resection mediated through Exonuclease 1.
Elongator-dependent modification of cytoplasmic tRNALysUUU is required for mitochondrial function under stress conditions
To gain a wider view of the pathways that regulate mitochondrial function, we combined the effect of heat stress on respiratory capacity with the discovery potential of a genome-wide screen in Saccharomyces cerevisiae. We identified 105 new genes whose deletion impairs respiratory growth at 37°C by interfering with processes such as transcriptional regulation, ubiquitination and cytosolic tRNA wobble uridine modification via 5-methoxycarbonylmethyl-2-thiouridine formation. The latter process, specifically required for efficient decoding of AA-ending codons under stress conditions, was covered by multiple genes belonging to the Elongator (e.g. ELP3) and urmylation (e.g., NCS6) pathways. ELP3 or NCS6 deletants had impaired mitochondrial protein synthesis. Their respiratory deficiency was selectively rescued by overexpression of tRNALysUUU as well by overexpression of genes (BCK1 and HFM1) with a strong bias for the AAA codon read by this tRNA. These data extend the mitochondrial regulome, demonstrate that heat stress can impair respiration by disturbing cytoplasmic translation of proteins critically involved in mitochondrial function and document, for the first time, the involvement in such process of the Elongator and urmylation pathways. Given the conservation of these pathways, the present findings may pave the way to a better understanding of the human mitochondrial regulome in health and disease.
Mechanism of CRISPR-RNA guided recognition of DNA targets in Escherichia coli
In bacteria and archaea, short fragments of foreign DNA are integrated into Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR) loci, providing a molecular memory of previous encounters with foreign genetic elements. In Escherichia coli, short CRISPR-derived RNAs are incorporated into a multi-subunit surveillance complex called Cascade (CRISPR-associated complex for antiviral defense). Recent structures of Cascade capture snapshots of this seahorse-shaped RNA-guided surveillance complex before and after binding to a DNA target. Here we determine a 3.2 Å x-ray crystal structure of Cascade in a new crystal form that provides insight into the mechanism of double-stranded DNA binding. Molecular dynamic simulations performed using available structures reveal functional roles for residues in the tail, backbone and belly subunits of Cascade that are critical for binding double-stranded DNA. Structural comparisons are used to make functional predictions and these predictions are tested in vivo and in vitro. Collectively, the results in this study reveal underlying mechanisms involved in target-induced conformational changes and highlight residues important in DNA binding and protospacer adjacent motif recognition.
The impact of the phosphomimetic eIF2{alpha}S/D on global translation, reinitiation and the integrated stress response is attenuated in N2a cells
A plethora of stresses trigger a rapid downregulation of protein synthesis. However, a fraction of mRNAs continue to be recruited onto polysomes and their protein products play a key role in deciding cell fate. These transcripts are characterized by the presence of uORFs within their 5' TL coupling protein expression to reinitiation. The translational brake arises due to the activation of a family of kinases targeting the α subunit of the trimolecular eIF2(αβ) initiation factor. Phosphorylation of eIF2αSer51 inhibits ternary complex regeneration reducing the pool of 43S ribosomes. It is popular to mimic this event, and hence the integrated stress response (ISR), by the expression of the phosphomimetic eIF2αS51D. However, we report that whereas the ISR is reproduced by eIF2αS51D expression in human HEK293T cells this is not the case in N2a mouse neuroblastoma cells. With regards to translational downregulation, this arises due to the failure of the phosphomimetic protein to assemble an eIF2 complex with endogenous eIF2β/. This can be compensated for by the transient co-expression of all three subunits. Curiously, these conditions do not modulate reinitiation and consequently fail to trigger the ISR. This is the first demonstration that the inhibitory and reinitiation functions of eIF2αS/D can be separated.
Competitive interaction of monovalent cations with DNA from 3D-RISM
The composition of the ion atmosphere surrounding nucleic acids affects their folding, condensation and binding to other molecules. It is thus of fundamental importance to gain predictive insight into the formation of the ion atmosphere and thermodynamic consequences when varying ionic conditions. An early step toward this goal is to benchmark computational models against quantitative experimental measurements. Herein, we test the ability of the three dimensional reference interaction site model (3D-RISM) to reproduce preferential interaction parameters determined from ion counting (IC) experiments for mixed alkali chlorides and dsDNA. Calculations agree well with experiment with slight deviations for salt concentrations >200 mM and capture the observed trend where the extent of cation accumulation around the DNA varies inversely with its ionic size. Ion distributions indicate that the smaller, more competitive cations accumulate to a greater extent near the phosphoryl groups, penetrating deeper into the grooves. In accord with experiment, calculated IC profiles do not vary with sequence, although the predicted ion distributions in the grooves are sequence and ion size dependent. Calculations on other nucleic acid conformations predict that the variation in linear charge density has a minor effect on the extent of cation competition.
Discovery of an essential nucleotidylating activity associated with a newly delineated conserved domain in the RNA polymerase-containing protein of all nidoviruses
RNA viruses encode an RNA-dependent RNA polymerase (RdRp) that catalyzes the synthesis of their RNA(s). In the case of positive-stranded RNA viruses belonging to the order Nidovirales, the RdRp resides in a replicase subunit that is unusually large. Bioinformatics analysis of this non-structural protein has now revealed a nidoviral signature domain (genetic marker) that is N-terminally adjacent to the RdRp and has no apparent homologs elsewhere. Based on its conservation profile, this domain is proposed to have nucleotidylation activity. We used recombinant non-structural protein 9 of the arterivirus equine arteritis virus (EAV) and different biochemical assays, including irreversible labeling with a GTP analog followed by a proteomics analysis, to demonstrate the manganese-dependent covalent binding of guanosine and uridine phosphates to a lysine/histidine residue. Most likely this was the invariant lysine of the newly identified domain, named nidovirus RdRp-associated nucleotidyltransferase (NiRAN), whose substitution with alanine severely diminished the described binding. Furthermore, this mutation crippled EAV and prevented the replication of severe acute respiratory syndrome coronavirus (SARS-CoV) in cell culture, indicating that NiRAN is essential for nidoviruses. Potential functions supported by NiRAN may include nucleic acid ligation, mRNA capping and protein-primed RNA synthesis, possibilities that remain to be explored in future studies.
Involvement of SRSF11 in cell cycle-specific recruitment of telomerase to telomeres at nuclear speckles
Telomerase, a unique ribonucleoprotein complex that contains the telomerase reverse transcriptase (TERT), the telomerase RNA component (TERC) and the TERC-binding protein dyskerin, is required for continued cell proliferation in stem cells and cancer cells. Here we identify SRSF11 as a novel TERC-binding protein that localizes to nuclear speckles, subnuclear structures that are enriched in pre-messenger RNA splicing factors. SRSF11 associates with active telomerase enzyme through an interaction with TERC and directs it to nuclear speckles specifically during S phase of the cell cycle. On the other hand, a subset of telomeres is shown to be constitutively present at nuclear speckles irrespective of cell cycle phase, suggesting that nuclear speckles could be the nuclear sites for telomerase recruitment to telomeres. SRSF11 also associates with telomeres through an interaction with TRF2, which facilitates translocation of telomerase to telomeres. Depletion of SRSF11 prevents telomerase from associating with nuclear speckles and disrupts telomerase recruitment to telomeres, thereby abrogating telomere elongation by telomerase. These findings suggest that SRSF11 acts as a nuclear speckle-targeting factor that is essential for telomerase association with telomeres through the interactions with TERC and TRF2, and provides a potential target for modulating telomerase activity in cancer.
A unique uracil-DNA binding protein of the uracil DNA glycosylase superfamily
Uracil DNA glycosylases (UDGs) are an important group of DNA repair enzymes, which pioneer the base excision repair pathway by recognizing and excising uracil from DNA. Based on two short conserved sequences (motifs A and B), UDGs have been classified into six families. Here we report a novel UDG, UdgX, from Mycobacterium smegmatis and other organisms. UdgX specifically recognizes uracil in DNA, forms a tight complex stable to sodium dodecyl sulphate, 2-mercaptoethanol, urea and heat treatment, and shows no detectable uracil excision. UdgX shares highest homology to family 4 UDGs possessing Fe-S cluster. UdgX possesses a conserved sequence, KRRIH, which forms a flexible loop playing an important role in its activity. Mutations of H in the KRRIH sequence to S, G, A or Q lead to gain of uracil excision activity in MsmUdgX, establishing it as a novel member of the UDG superfamily. Our observations suggest that UdgX marks the uracil-DNA for its repair by a RecA dependent process. Finally, we observed that the tight binding activity of UdgX is useful in detecting uracils in the genomes.
Post-transcriptional gene silencing triggered by sense transgenes involves uncapped antisense RNA and differs from silencing intentionally triggered by antisense transgenes
Although post-transcriptional gene silencing (PTGS) has been studied for more than a decade, there is still a gap in our understanding of how de novo silencing is initiated against genetic elements that are not supposed to produce double-stranded (ds)RNA. Given the pervasive transcription occurring throughout eukaryote genomes, we tested the hypothesis that unintended transcription could produce antisense (as)RNA molecules that participate to the initiation of PTGS triggered by sense transgenes (S-PTGS). Our results reveal a higher level of asRNA in Arabidopsis thaliana lines that spontaneously trigger S-PTGS than in lines that do not. However, PTGS triggered by antisense transgenes (AS-PTGS) differs from S-PTGS. In particular, a hypomorphic ago1 mutation that suppresses S-PTGS prevents the degradation of asRNA but not sense RNA during AS-PTGS, suggesting a different treatment of coding and non-coding RNA by AGO1, likely because of AGO1 association to polysomes. Moreover, the intended asRNA produced during AS-PTGS is capped whereas the asRNA produced during S-PTGS derives from 3' maturation of a read-through transcript and is uncapped. Thus, we propose that uncapped asRNA corresponds to the aberrant RNA molecule that is converted to dsRNA by RNA-DEPENDENT RNA POLYMERASE 6 in siRNA-bodies to initiate S-PTGS, whereas capped asRNA must anneal with sense RNA to produce dsRNA that initiate AS-PTGS.